A Practical Guide to Family Studies with Lifetime Data
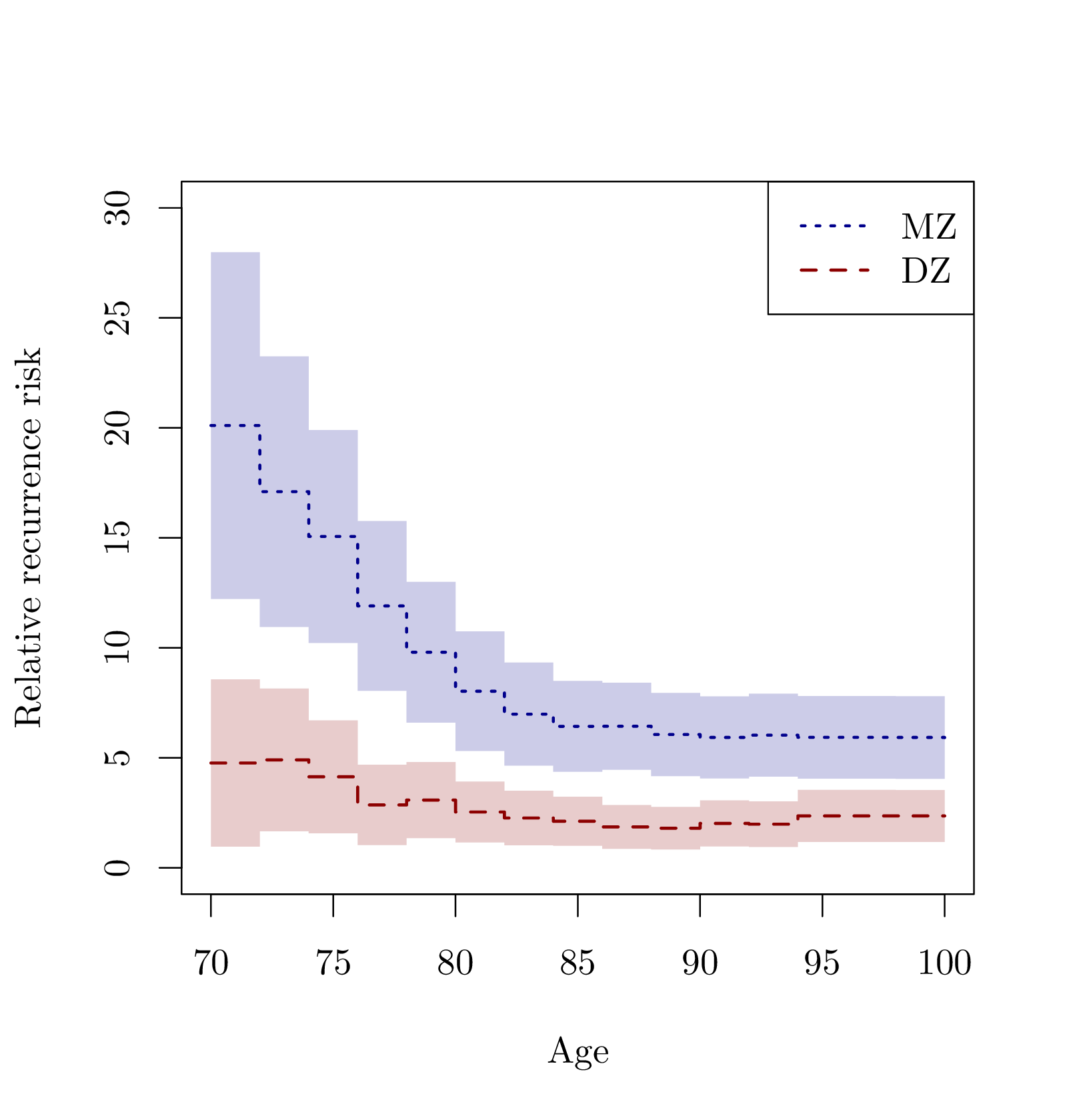
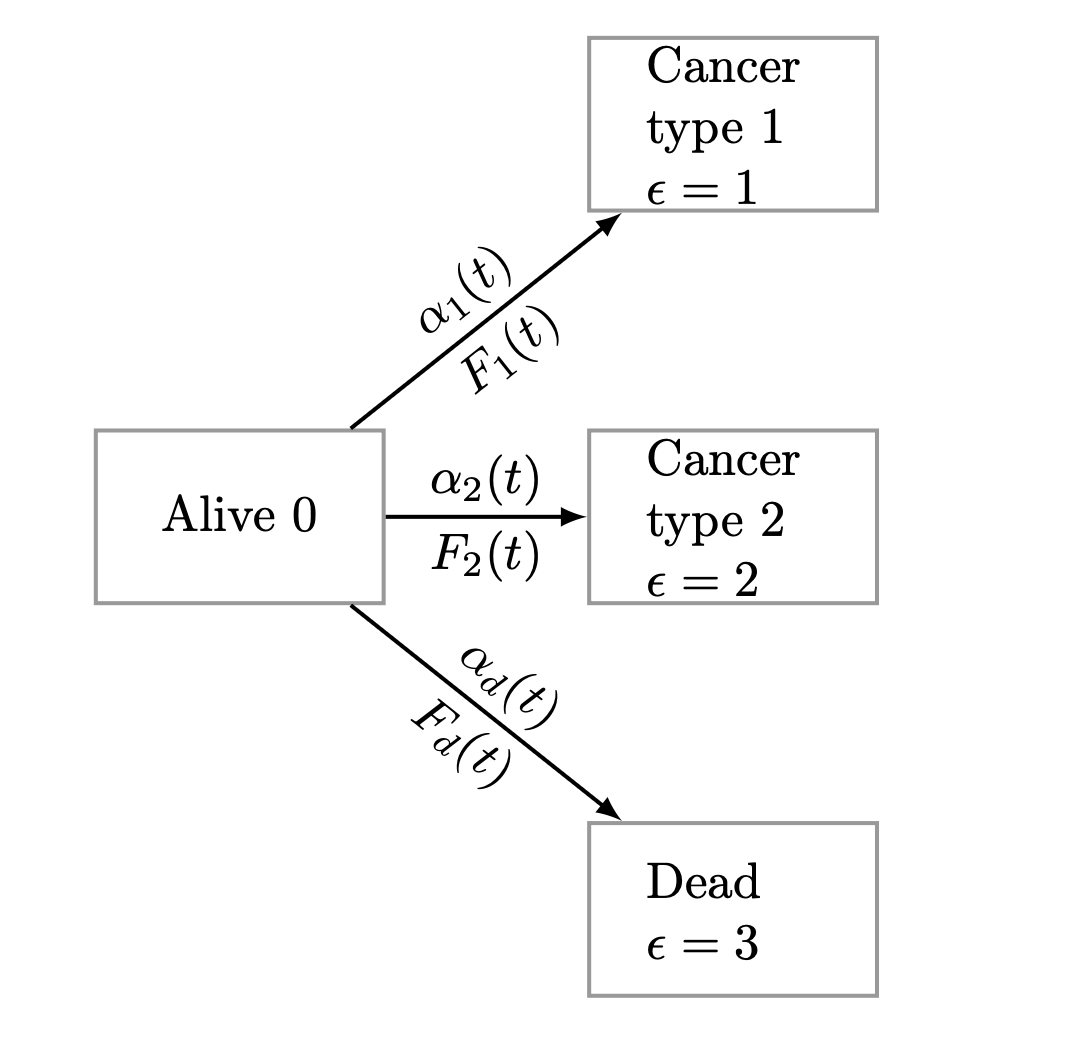
Familial aggregation refers to the fact that a particular disease may be overrepresented in some families due to genetic or environmental factors. When studying such phenomena, it is clear that one important aspect is the age of onset of the disease in question, and in addition, the data will typically be right-censored. Therefore, one must apply lifetime data methods to quantify such dependence and to separate it into different sources using polygenic modeling. Another important point is that the occurrence of a particular disease can be prevented by death mdash that is, competing risks mdash and therefore, the familial aggregation should be studied in a model that allows for both death and the occurrence of the disease. We here demonstrate how polygenic modeling can be done for both survival data and competing risks data dealing with right-censoring. The competing risks modeling that we focus on is closely related to the liability threshold model. doi: “10.48550/arXiv.2502.03942”